Today I updated the RNA isolation protocol based on info from Sam, and I made some mistakes in isolating RNA today. Ran Qubit.
Updated RNA isolation protocol for crab hemolymph to be used starting tomorrow. Link: here
RNA Isolation
I had previously thought that RNAzol was a dye that binded to RNA… I double-checked with Sam, because I realized I never really asked, and after centrifuging, the RNAzol (blue) is often at the top of the tube, but sometimes it’s in the middle or towards the bottom and I wasn’t sure if I should be aiming for the blue when removing the supernatant for further processing. He said the RNAzol is actually salts and other stuff that helps RNA become isolated. So, when taking out the supernatant, it’s ok if I get some blue, but I should aim for the clear liquid, while avoiding the bottom, becasue that’s where the junk gets pelleted down to.
Today I ran 9 more samples:
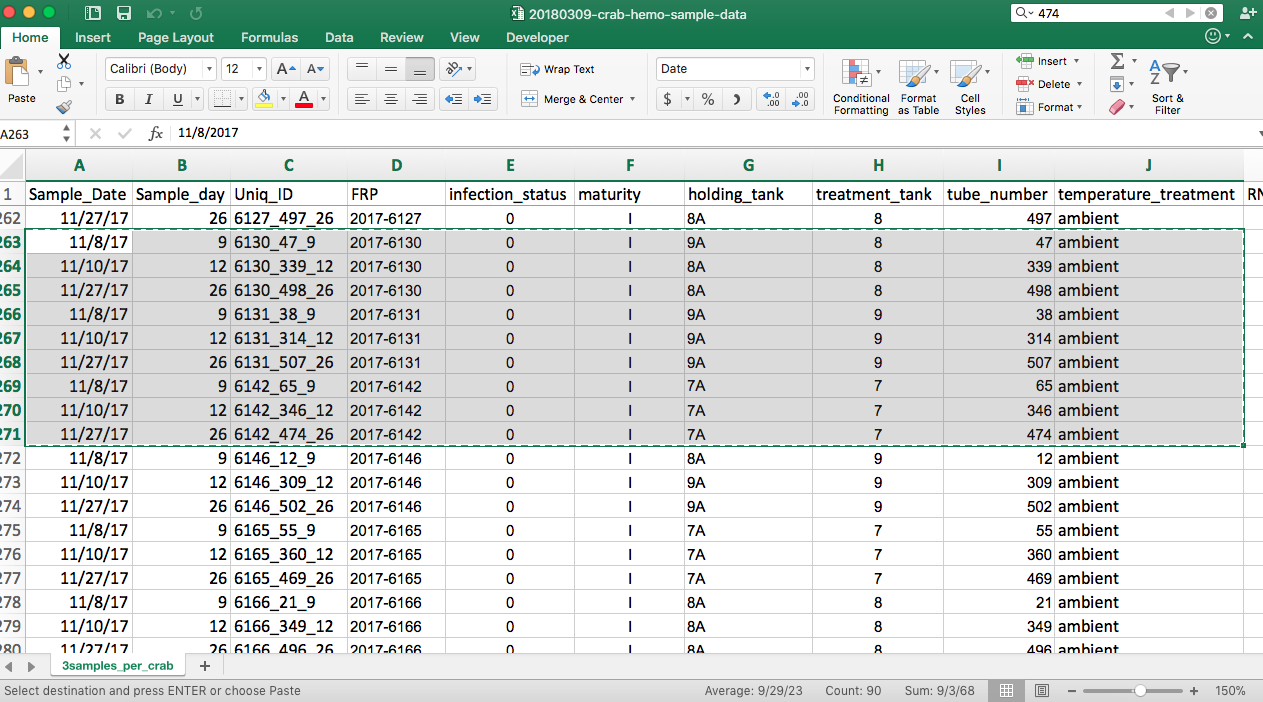
I had a couple of mishaps today :( Firstly, during the step where I remove the supernatant into a new tube, I accidentally put the tip of the pipet too far down, and I think I likely got some junk in with the supernatant for tube # 474. Luckily tube 474 is from the third hemolymph sampling day, which means that I have two more samples from that same crab for that day.
Secondly, when adding 750µL of 2-propanol, I accidentally tipped tube #38 (first sampling day) and lost some of the supernatant that I had saved.
Here is Sam’s notebook that I should have looked at wayyyy before starting this process. I had forgotten that he did this on two crab samples a while back. link
I see that Sam mixed the 2-propanol with the supernatant by inversion rather than vortex. I think I will start doing this instead, because the vortex is so powerful that the propanol often ends up leaking a little, and smearing the tube labels.
Qubit results
Used 5µL of sample from tubes 65 and 498. 65 was out of range, 498 was 1.8ng/µL.
